Fig.1 Homepage of the Bitter Gourd Resource Database
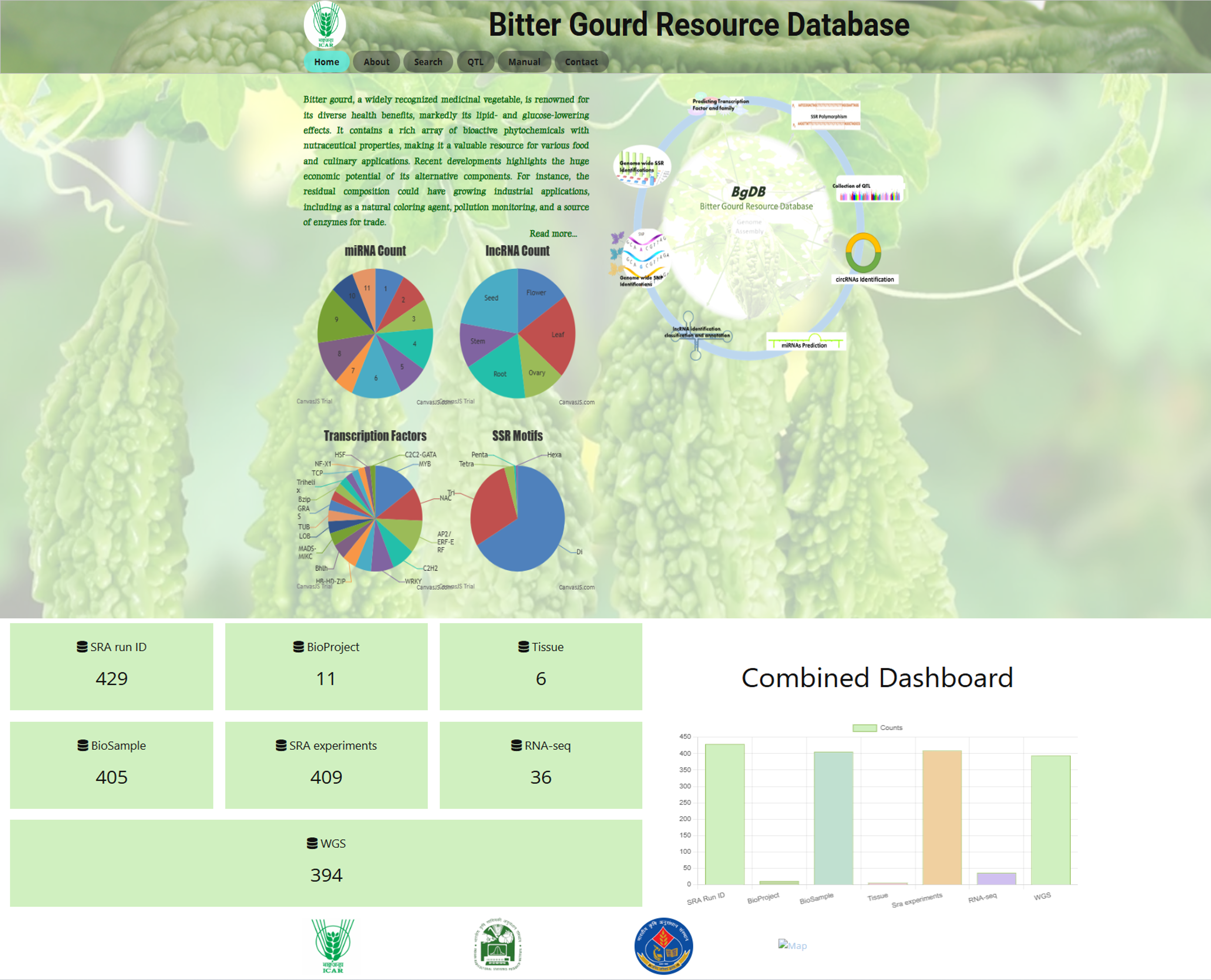
The Homepage provides key sections for easy navigation:
Fig.2 About Section of the Bitter Gourd Resource Database

The About section of the Bitter Gourd Resource Database provides an introduction to the database and its focus on bitter gourd (Momordica charantia L.), a significant vegetable crop in the Cucurbitaceae family.
Fig.3 Search Section of the Bitter Gourd Resource Database

The Search section of the Bitter Gourd Resource Database allows users to query and retrieve specific genomic data related to bitter gourd (Momordica charantia L.). This section is designed to be user-friendly and provides various options to filter and refine search results.
The Variant section in the Search tool allows users to explore genetic variations in bitter gourd. It includes three types:
Fig.4 Variant Section of the Bitter Gourd Resource Database

Fig.5 SSR section of the Bitter Gourd Resource Database
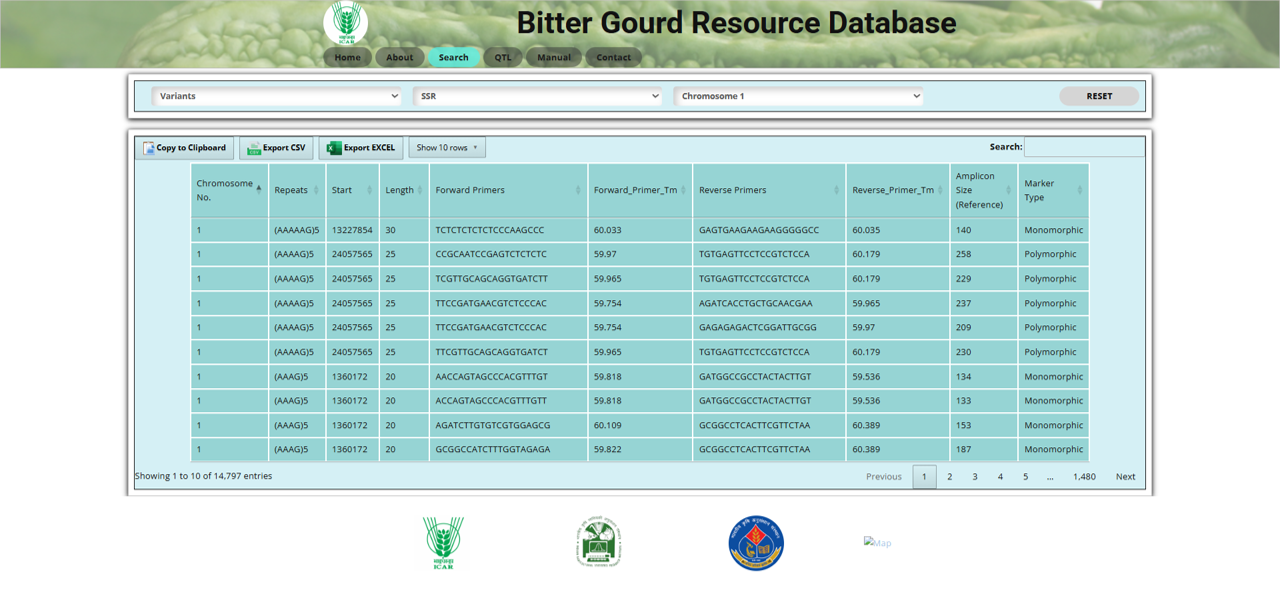
Fig.6 SNP section of the Bitter Gourd Resource Database
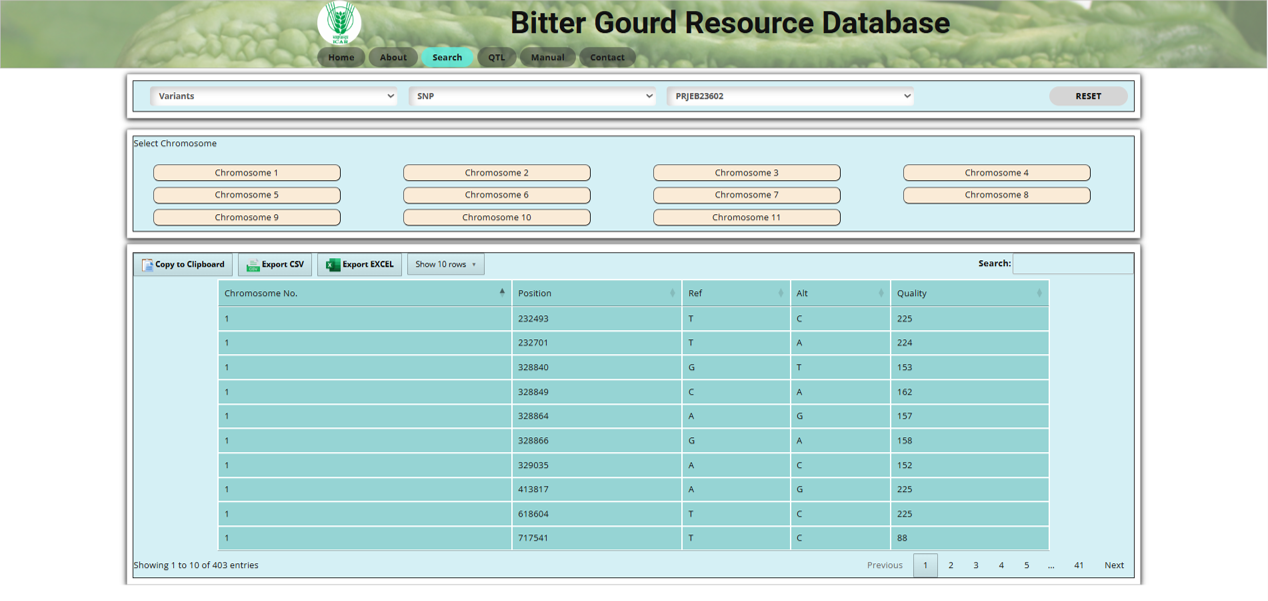
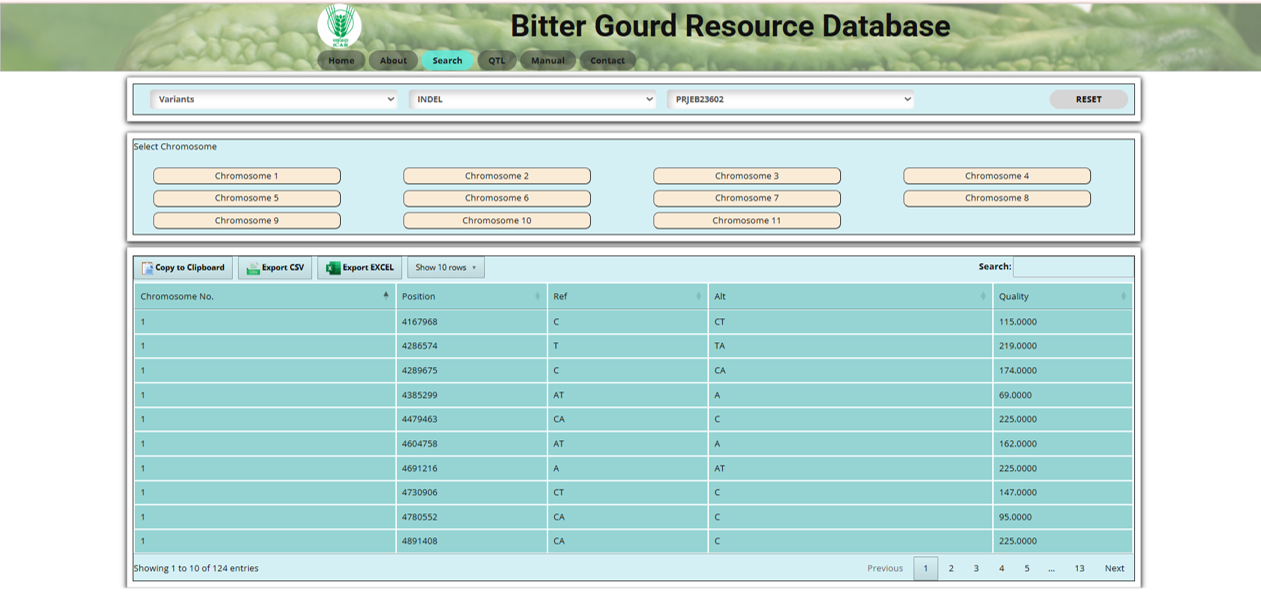
Fig.7 INDEL section of the Bitter Gourd Resource Database
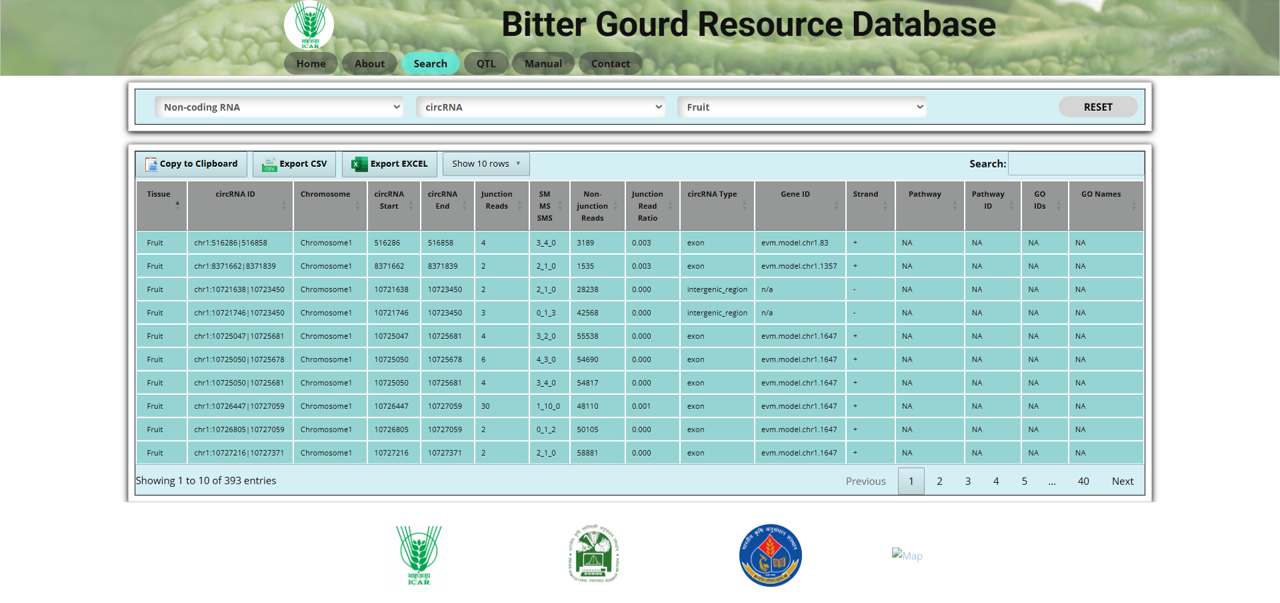
The Non-coding RNA section allows users to explore different types of non-coding RNAs in bitter gourd (Momordica charantia L.), which play crucial roles in gene regulation and cellular processes. The section includes three options:
In the circRNA section, users can:
Fig.8 circRNA section of the Bitter Gourd Resource Database
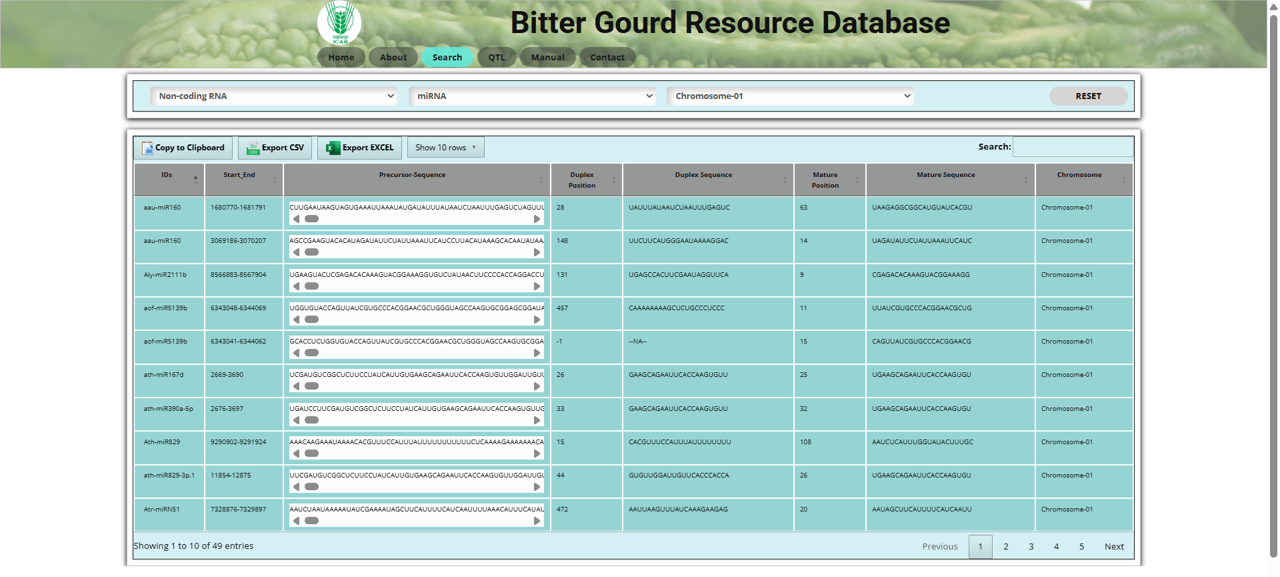
The miRNA section in the Non-coding RNA search tool allows users to explore microRNAs (miRNAs) in bitter gourd (Momordica charantia L.). miRNAs are small non-coding RNAs that play a crucial role in gene regulation.
In the miRNA section, users can:
The lncRNA section in the Non-coding RNA search tool allows users to explore long non-coding RNAs (lncRNAs) in bitter gourd (Momordica charantia L.). lncRNAs are involved in various regulatory processes within the cell.
In the lncRNA section, users can:
Fig.9 lncRNA section of the Bitter Gourd Resource Database
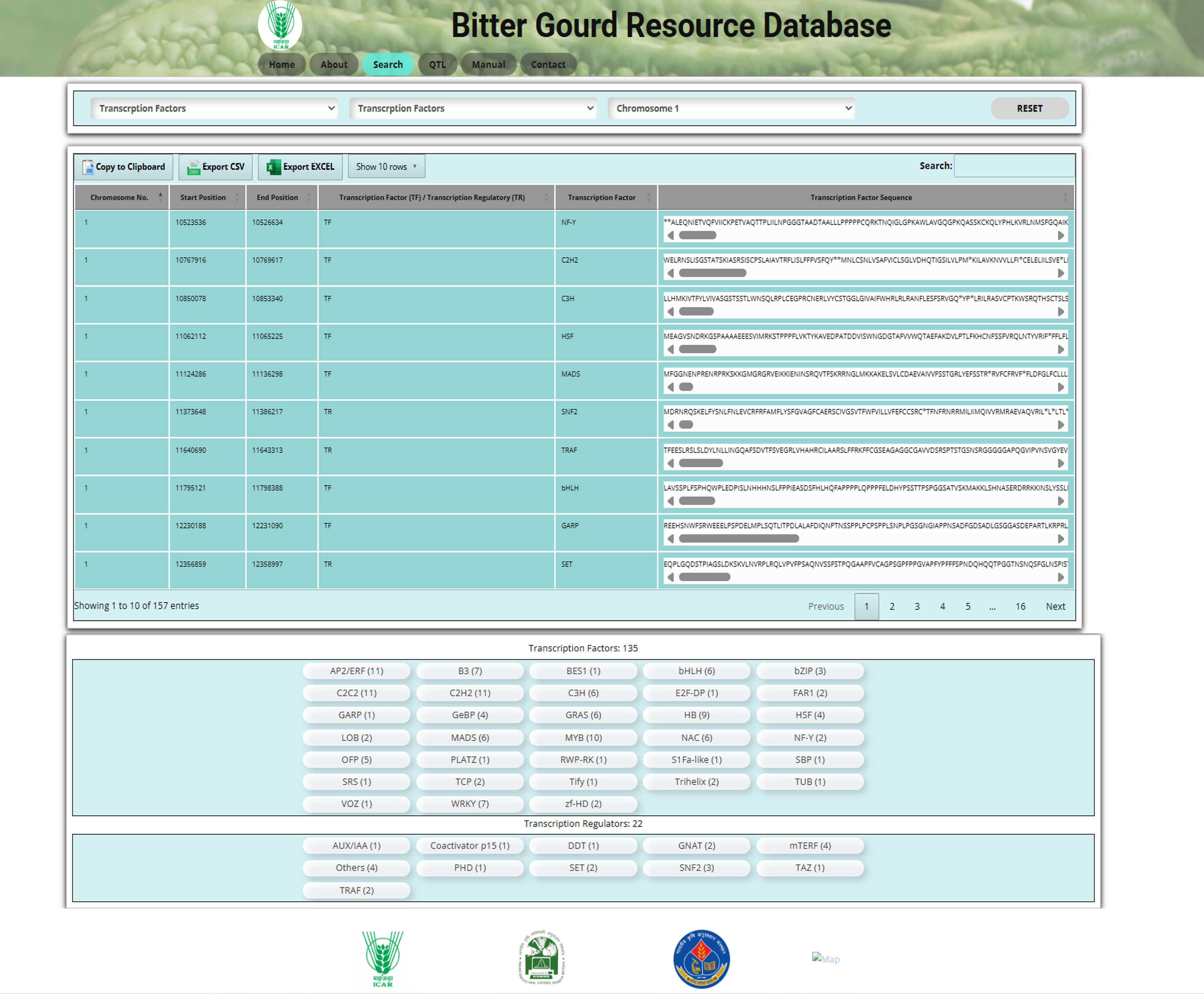
The Transcription Factors section in the Search tool allows users to explore transcription factors in bitter gourd (Momordica charantia L.). Transcription factors are proteins that regulate gene expression by binding to specific DNA sequences.
Fig.10 Transcription factor section of the Bitter Gourd Resource Database
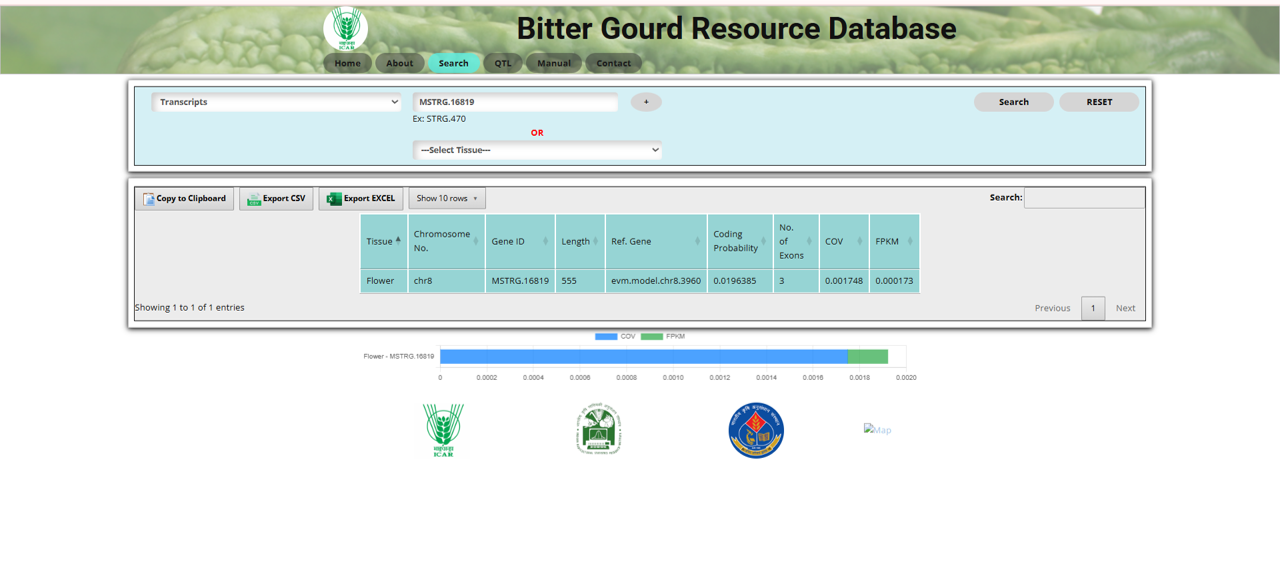
The Transcripts section in the Search tool allows users to explore transcript data in bitter gourd (Momordica charantia L.). Transcripts represent the RNA molecules produced from the transcription of DNA and are crucial for understanding gene expression.
Fig.11 Transcript section of the Bitter Gourd Resource Database
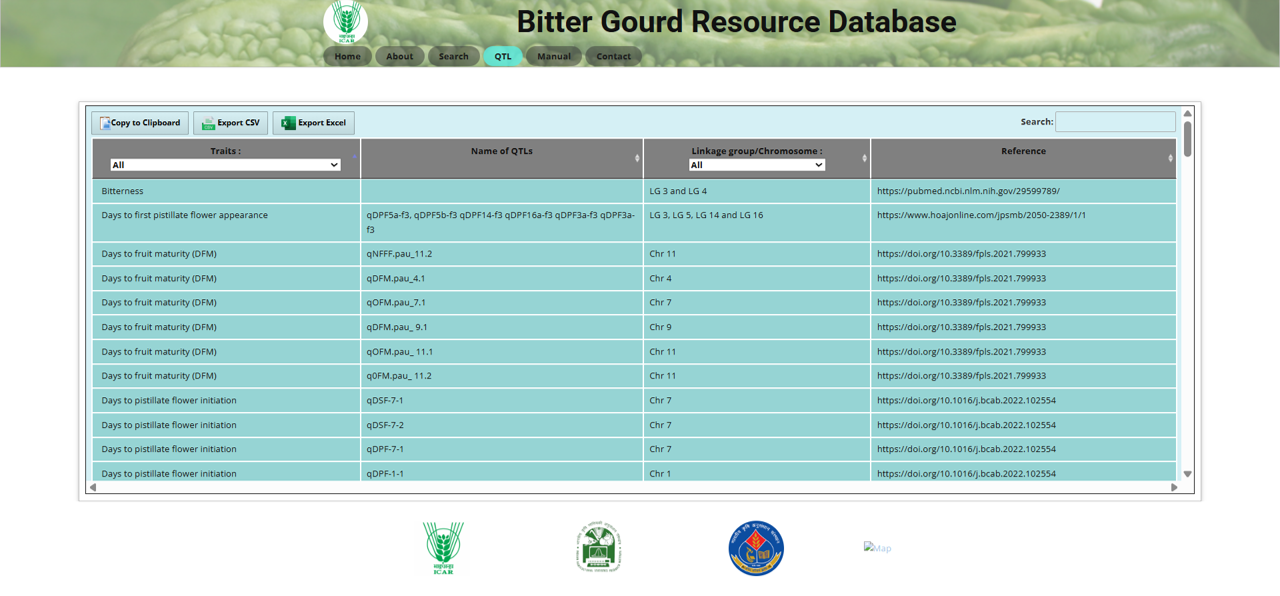
The QTL (Quantitative Trait Loci) section allows users to explore genetic loci associated with traits in bitter gourd. Key features include:
Fig.13 QTL section of the Bitter Gourd Resource Database
The Contact page provides information for reaching out to the team behind the Bitter Gourd Resource Database.
Fig.14 Contact section of the Bitter Gourd Resource Database